Citizen scientists, AI and cloud bond to boost billabong health in the Top End
A major initiative is underway in Australia’s Top End – combining citizen scientists, artificial intelligence and cloud computing to create the world’s largest training dataset of fish species.
Armed with that dataset it will be far easier to train machine learning models to correctly identify different species of fish; a critical input to assessing and monitoring global fish stocks.
The Australian Government’s Supervising Scientist Branch (SSB) is leading the initiative.
The SSB is tasked with protecting the dual-world-heritage listed Kakadu National Park from the effects of uranium mining. It does this by undertaking environmental research and monitoring to develop standards and practices that ensure environmental protection.
Ultimately, the SSB wants to develop innovative, easily deployable monitoring tools that can be used by Indigenous Ranger groups to undertake the long-term monitoring of the rehabilitated Ranger uranium mine site.
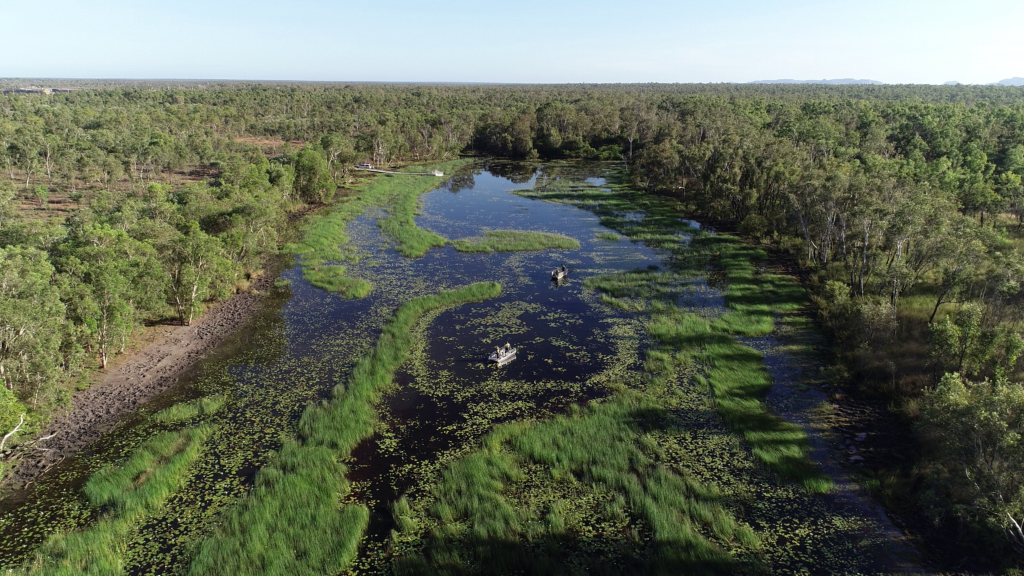
One way it measures the health of aquatic ecosystems is through monitoring changes in fish community composition in channel and shallow lowland billabongs up and downstream of the Ranger uranium mine in the Northern Territory.
This was previously tackled by teams of up to 15 people working in the field for weeks at a time, using nets to sample fish or counting them by peering through see-through panes in the hull of the SSB’s bubble boat.
However increased crocodile activity in the area made that a perilous proposition, and the SSB instead began collecting videos from underwater cameras and drones. While scientists were kept safe from crocodile attacks – they risked drowning in the deluge of data that needed to be logged and analysed.
Scientists learned in 2018 of an AI-based solution developed to identify fish in Darwin Harbour that was the result of a partnership between the NT Department of Primary Industry and Resources (DPIR) fisheries team and Microsoft.
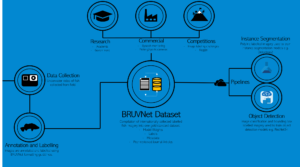
The BRUVNet (baited remote underwater video net) that SSB is working on is an open sourced set of fish images that can be used to train machine learning platforms to automatically identify fish species from video.

Could that also be applied to monitor fish in the Territory’s billabongs?
To learn the answer to that question Microsoft spoke with Andrew Jansen, Bio-monitoring Manager, Water and Sediment Quality, from DAWE.
Microsoft (MS): What led you to this platform?
Andrew Jansen (AJ): Seeing the media pieces from Darwin Harbour immediately grabbed our attention as we were seeking to replicate the same automation techniques for our freshwater fish species found in Kakadu National Park. This initiated our engagement with Microsoft’s machine learning and data science FastTrack for Azure Lead Engineer, Steve van Bodegraven, who began educating us on all the learnings from NT Fisheries, and led us on the path to where we are now.
By partnering with Microsoft and using AI and Deep Learning to automate the data processing and analyses in the back end, we will have a highly complex and scientifically rigorous monitoring tool that can be easily deployed in the field.
MS: What impact are you hoping to achieve?
AJ: We will generate the largest training dataset of fish species in the world. We want to enable all fisheries scientists to use cloud computing and artificial intelligence to automate what is now a very manual process in quantifying fish stocks around the globe. Fisheries scientists are drowning in BRUV footage so this technology will allow greater spatial and temporal sampling of fish and increase our understanding of fish populations on a global scale.
For the Supervising Scientist Branch, when fish ecologists no longer need to manually process fish videos it frees up time for other research. Nationally, we’re working towards greater collaboration between fisheries scientists to share learnings, datasets and models, and to see fish monitoring programs using the pipelines developed here to automate processing of their own videos.
MS: How does this work?
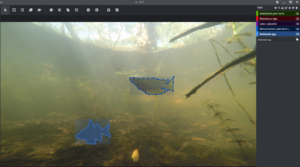
AJ: Our underwater cameras (GoPros) are deployed on surface floating and bottom-dwelling stands along the bank of the channel billabongs pointing toward habitat for fish species and amongst aquatic vegetation in our shallow lowland billabongs.
The video from that was used with CustomVision.ai for image labelling using bounding boxes and for training AI models. This allowed us to understand how our data performed in a really quick way – a lot of learnings came out of using this tool.
We also used Azure Batch to scale compute power across 20+ Virtual Machines and speed up data processing. And now, Azure Machine Learning service is used to train the BRUVNet dataset on the latest AI models to segment fish in the images to species level.
PowerBI is used to visualise the BRUVNet dataset status (number of species, annotations, labels, images etc.), and to derive our fish community abundance metrics from scored videos.


MS: How are citizen scientists helping?
AJ: On our journey, we learnt that polygon labelled imagery will return the best result when trying to identify fish to species level using deep learning. Using CustomVision.ai we were able to develop a bounding box labelled dataset relatively quickly using fish ecologists in the NT, however due to the complexity of the data, polygons are required. This takes a significant amount of time to trace the outline of all fish species in an image, that government and research institutes can’t adequately fund to achieve the desired model performance. Engaging citizen scientists to perform the fish tracing is one way of speeding up the annotation process.
MS: How do you train citizen scientists to do this?
AJ: We have used all open-source free tools to enable anyone to replicate our process and make it as simple as possible. As we are only asking people to trace lines around fish – applying species tags is an optional bonus – the process is actually quite simple and allows anyone to open VoTT (Visual Object Tagging Tool) in a web browser, enter some unique codes to connect to images stored in Azure Cloud Containers and using our one-page instruction guide start tracing lines around the fish in their images.
All data and labels are automatically saved in the cloud which means no data is lost and there is no requirement to save or backup work that is completed. We provide a photo guide for those who want to try their hand at tagging fish with species name, however the Supervising Scientists’ fish ecologists will be checking the quality of the annotations and species tags. Whilst this represents a significant workload, it will ensure the BRUVNet dataset is a true gold standard that can be used by all fisheries scientists.
MS: How do you envisage this being used?
AJ: If successful, the image labelling challenge will generate the largest international dataset of polyline labelled freshwater fish imagery that meets quality control requirements. It will be used to train an AI model that will automate the Supervising Scientists’ fish video processing saving hundreds of hours of manual work each year. As the dataset and model weights will be made publicly available we hope it will encourage other fisheries scientists working in this space to contribute their data, and those who want to implement this technology will be able to fast track this as a routine monitoring tool to assess ecosystem health.
MS: When do you plan to go live?
AJ: We have undergone a soft launch within the Department to test and improve the processes. This is now complete. We have all the notebooks and draft GitHub Repository at 90 per cent, all that is required is building into an Azure Template so when we go live others can immediately use what we’ve achieved and deploy in Azure. We aim for this release in early October.
MS: What’s next on your agenda?
AJ: The Supervising Scientist runs a drone program which is used to characterise the surrounding landscape (terrestrial ecosystem) that will be used to inform rehabilitation of Ranger uranium mine. We’re looking at adopting some of the skills and learnings from BRUVNet to automate processing of our Lidar, multi-spectral and hyperspectral imagery to classify tree, understory and aquatic vegetation species.